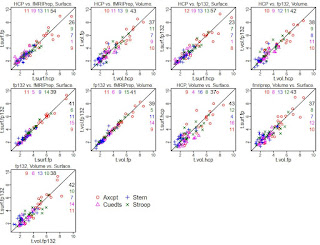
In this post I show the results of correlating the functional connectivity matrices, on the suggestion of Max Bertolero (@max_bertolero, thanks!). Specifically, I turned the lower triangles of each connectivity matrix into a vector then correlated them pairwise. (In other words, "unwrapping" each functional connectivity matrix into a vector then calculating all possible pairwise correlations between the 26 vectors.)
The results are in the figure below. To walk you through it: participants are along the x-axis, matrix-to-matrix correlations on the y-axis. Each person is given a unique plotting symbol, which is shown in black. The first column is for person 150423, whose plotting symbol is a circle. The correlation of 150423's hcp-Siegel and fmriprep-xcp matrices is about 0.75 (location of the black circle). The four colored sets of symbols are the correlation of 150423's hcp or fmriprep matrix with everyone else's hcp or fmriprep matrices, as listed along the top. For example, the bright blue column is listed as "hcp w/others' fp", and the highest symbol is a plus. Looking at the black symbols, the plus sign is 171330, so this means that the correlation of 150423's hcp-Siegel matrix with 171330's fmriprep-xcp matrix is about 0.63.
There are several interesting parts of the figure. First, the dark blue and dark red points tend to be a bit higher than the bright blue and pink: matrices derived from the same pipeline tend to have higher correlation than those from different pipelines, even from different people.
Next, most people's fmriprep and hcp matrices are more correlated than either are to anyone else's - the black symbols are usually above all of the colored ones. This is not the case for 3 people: 178950, DMCC8033964, and DMCC9478705. 203418 is also interesting, since their self-correlation is a bit lower than most, but still higher than the correlation with all the others.
Finally, I compared the impressions from this graph with those from the blinded matching (summarized in the table below), and it came out fairly consistently, particularly for the hardest to match pairs: No one correctly matched 178950 or DMCC9478705's matrices, two of the people just listed as having their self-correlations in the midst of the others. Only one person correctly matched the third (DMCC8033964).
While the hardest-to-match pairs are easy to see in this graph, the ones most often correctly matched are more difficult to predict. For example, from the height of the self-correlation and distance to the other points I'd predict that 171330 and 393550 would be relatively hard to pair, but they were the most often correctly matched. 203418 has the fourth-lowest self-correlation, but was correctly matched by more than half the testers.
Note: the "correct matches" column is how many of the 9 people doing the blinded matching test correctly matched the person's matrices.
| subject ID | correct pairing (hcp, fmriprep) | correlation of this person’s matrices | correct matches |
| 150423 | h,q | 0.7518 | 4 |
| 155938 | v,i | 0.7256 | 6 |
| 171330 | l,y | 0.7643 | 4 |
| 178950 | b,r | 0.6069 | 0 |
| 203418 | x,d | 0.651 | 5 |
| 346945 | w,j | 0.7553 | 4 |
| 393550 | u,n | 0.7772 | 7 |
| 601127 | c,m | 0.7152 | 1 |
| 849971 | k,t | 0.7269 | 1 |
| DMCC5775387 | s,f | 0.767 | 6 |
| DMCC6705371 | a,p | 0.733 | 6 |
| DMCC8033964 | g,z | 0.5938 | 1 |
| DMCC9478705 | o,e | 0.5428 | 0 |
Together, both the blinded matching and correlation results reassure me that the two pipelines are giving qualitatively similar matrices, but clearly not identical, nor equally similar for all of the test people.
UPDATE 4 December 2019: Checking further, missings have a sensible impact on these results. These functional connectivity matrices were not calculated from the DMCC baseline session only (which are the runs included in DMCC13benchmark), but rather by concatenating all resting state runs for the person in the scanning wave. Concatenating gives 30 minutes of resting state total without missings or censoring.
Two subjects had missing resting state runs: DMCC8033964 is missing one (25 minutes total instead of 30), and DMCC9478705 is missing two (20 minutes). These are two of the lowest-correlation people, which is consistent with the expectation that longer scans make more stable functional connectivity matrices. It looks like the two pipelines diverge more when there's fewer input images, which isn't too surprising. 178950 didn't have missing runs but perhaps had more censoring or something; I don't immediately know how to extract those numbers from the output but expect the pipelines to vary in many details.
UPDATE 4 January 2021: Corrected DMCC13benchmark openneuro links.